Abstract
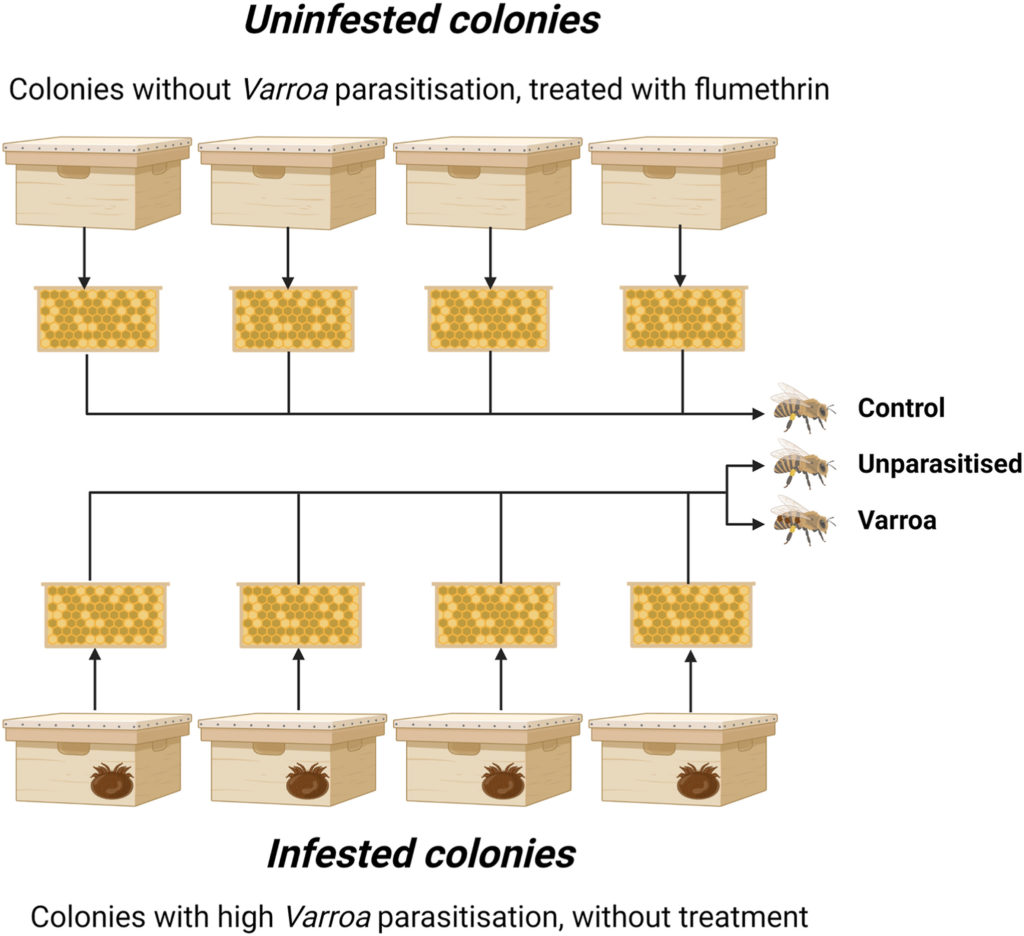
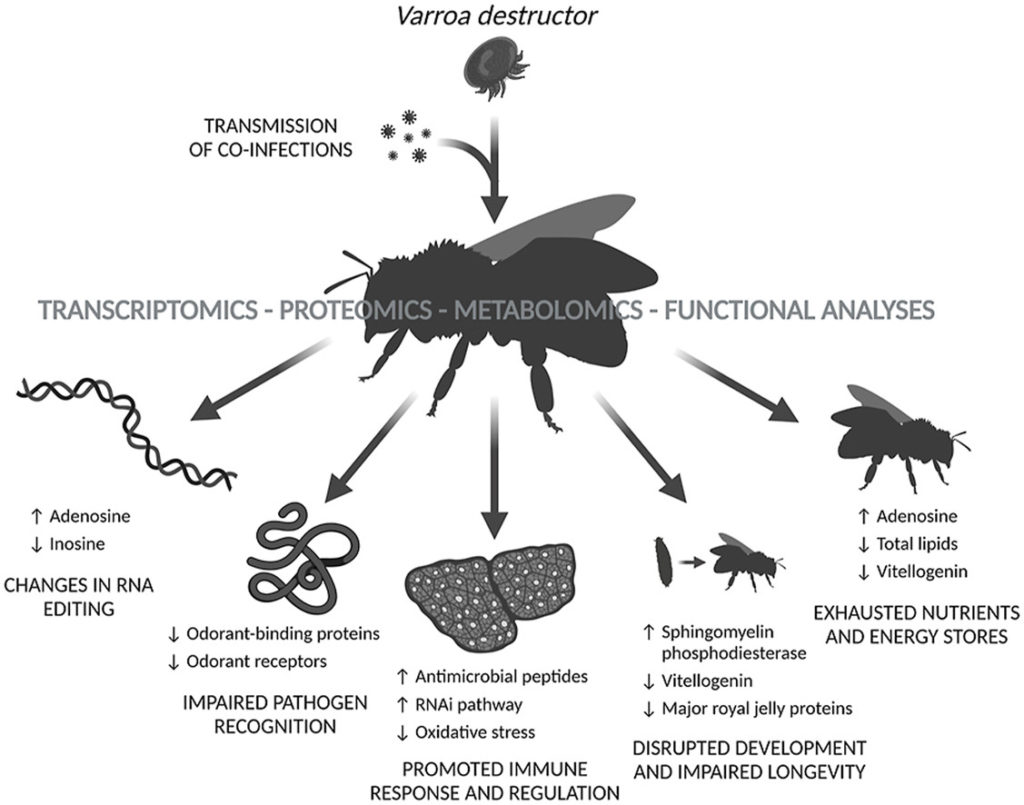
The extensive annual loss of honey bees (Apis mellifera L.) represents a global problem affecting agriculture and biodiversity. The parasitic mite Varroa destructor, associated with viral co-infections, plays a key role in this loss. Despite years of intensive research, the complex mechanisms of Varroa – honey bee interaction are still not fully defined. Therefore, this study employed a unique combination of transcriptomic, proteomic, metabolomic, and functional analyses to reveal new details about the effect of Varroa mites and naturally associated factors, including viruses, on honey bees. We focused on the differences between Varroa parasitised and unparasitised ten-day-old worker bees collected before overwintering from the same set of colonies reared without anti-mite treatment. Supplementary comparison to honey bees collected from colonies with standard anti-Varroa treatment can provide further insights into the effect of a pyrethroid flumethrin. Analysis of the honey bees exposed to mite parasitisation revealed alterations in the transcriptome and proteome related to immunity, oxidative stress, olfactory recognition, metabolism of sphingolipids, and RNA regulatory mechanisms. The immune response and sphingolipid metabolism were strongly activated, whereas olfactory recognition and oxidative stress pathways were inhibited in Varroa parasitised honey bees compared to unparasitised ones. Moreover, metabolomic analysis confirmed the depletion of nutrients and energy stores, resulting in a generally disrupted metabolism in the parasitised workers. The combined omics-based analysis conducted on strictly parasitised bees revealed the key molecular components and mechanisms underlying the detrimental effects of Varroa sp. and its associated pathogens. This study provides the theoretical basis and interlinked datasets for further research on honey bee response to biological threats and the development of efficient control strategies against Varroa mites.
Insect Biochem Mol Biol. 2022;103877. doi: 10.1016/j.ibmb.2022.103877.
Authors:
Martin Kunca, Pavel Dobeša*, Rachel Wardb, Saetbyeol Leec, Radim Čegand, Silvie Dostálkováe, Kateřina Holušováf, Jana Hurychováa, Sara Eliáša, Eliška Pinďákováe, Eliška Čukanovág, Jana Prodělalovág, Marek Petřivalskýe, Jiří Danihlíke, Jaroslav Havlíkc, Roman Hobzad, Kevin Kavanaghb, Pavel Hyršla
a Department of Experimental Biology, Faculty of Science, Masaryk University, Kamenice 5, 625 00, Brno, Czech Republic, b Department of Biology, Maynooth University, W23 F2K8 Maynooth, Co. Kildare, Ireland, c Department of Food Science, Faculty of Agrobiology, Food and Natural Resources, Czech University of Life Sciences Prague, Kamýcká 129, 165 00, Prague, Czech Republic, d Department of Plant Developmental Genetics, Institute of Biophysics of the Czech Academy of Sciences, Královopolská 135, 612 00, Brno, Czech Republic, e Department of Biochemistry, Faculty of Science, Palacký University Olomouc, Šlechtitelů 27, 783 71, Olomouc, Czech Republic, f Institute of Experimental Botany of the Czech Academy of Sciences, Centre of the Region Haná for Biotechnological and Agricultural Research, Šlechtitelů 31, 779 00, Olomouc, Czech Republic, g Department of Infectious Disease and Preventive Medicine, Veterinary Research Institute, Hudcova 296/70, 621 00, Brno, Czech Republic